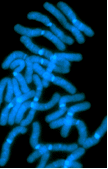
Metaphase chromosomes of slash pine (Pinus elliottii) stained with DAPI (see publication list and search with the keyword Pinus for details of our publications on pine; we also have collaborations with Dr Robert Doudrick, USDA Forest Service).
1. OBJECTIVES
1.1 Objectives The main objective of the proposed research is to improve our basic understanding of genome organisation in conifers. This will be achieved by developing and utilising a series of innovative, generic enabling technologies that will be used to characterise conifer genomes at various tiers of organisational complexity. Emphasis will be given to the following objectives: a. An examination of the content and distribution of repetitive DNA sequences to identify the location of such sequences within specific chromosomes and their conservation across species of the Pinaceae family. b. Determine level of synteny or conservation of gene order between species within the Pinaceae family. c. Characterisation and mapping of expressed sequence tags or EST's to serve as anchored sequences tagged sites (STS). d. Construction of comprehensive, informative second generation linkage maps for three Conifer genomes. e. Provide a sustainable biological resource to investigate the genetical control of quantitative traits in Conifers.
1.2 Compliance of the Objectives with the Priorities of the Work Programme a. Integrating genetic maps and models. Fluorescent in situ hybridisation (FISH) on metaphase chromosomes and on extended chromatin will be used to physically locate repetitive as well as single copy DNA sequences on conifer chromosomes and to integrate physical and genetic maps. b. Conifers are diploids which exhibit conservation of chromosome number (2n=2x=24). Initial studies with isozymes (1) indicate that pines exhibit conservation of gene order and this hypothesis will be examined further by the use of transcript based linkage maps. c. Single pass sequencing of cDNA clones provides a rapid route to gene discovery and production of EST's (2). Systematic sequencing of cDNA clones derived from xylem and leaf tissue will produce a new generation of research tools for conifer biology. d. Considerable progress has been achieved in Conifer genome mapping by the use of megagametophyte tissue and RAPDs (3). However, the utility of such maps is limited and there is a recognised need to produce second generation (STS based) linkage maps represented by living pedigrees. Segregating populations will be characterised with both single locus (microsatellites) and multi-locus (AFLPs) marker systems providing framework maps for the eventual location of cDNAs generated from single pass sequencing.
e. The establishment of synteny maps will provide a powerful tool to investigate the quantitative conservation of map location of trait loci across species and environments. The map position of cDNA clones relative to the predicted location of quantitative trait loci will provide new opportunities to unravel the genetic control of polygenic traits in forest tree species.