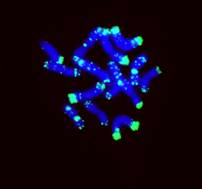
 Left: The distribution of the repetitive DNA sequence dpTa1
(DNA most diverse on D genome) using florescent in situ hybridization on the
chromosomes (stained blue with DAPI) in accession At524 of Aegilops tauschii.
Left: The distribution of the repetitive DNA sequence dpTa1
(DNA most diverse on D genome) using florescent in situ hybridization on the
chromosomes (stained blue with DAPI) in accession At524 of Aegilops tauschii.Diversity in Iranian wheats, particularly Triticum (Aegilops) tauschii (squarrosa; D genome)
Dr Hojjatollah Saeidi and Professor Mohammad Reza Rahiminejad, Isfahan University, Iran and
Pat Heslop-Harrison and Trude Schwarzacher, University of Leicester
 Left: The distribution of the repetitive DNA sequence dpTa1
(DNA most diverse on D genome) using florescent in situ hybridization on the
chromosomes (stained blue with DAPI) in accession At524 of Aegilops tauschii.
Left: The distribution of the repetitive DNA sequence dpTa1
(DNA most diverse on D genome) using florescent in situ hybridization on the
chromosomes (stained blue with DAPI) in accession At524 of Aegilops tauschii.
Iran, as a part of the Fertile Crescent, is within the centre
of diversity and centre of origin of most of Triticeae plants. Thus, the study
of biodiversity of this group within this area is important, giving information
about the range of genes present, origin of species, and origin of the polyploid species such as bread wheat. Because of the contribution of the D genome
to bread wheat, the genetic diversity present within Ae.
tauschii is of considerable interest for finding
new genes for exploitation in agriculture, and for tracking the origins of crop
domestication. It has been reported that the T. aestivum D-genome
originated from the SE or SW Caspian Sea in
We are exploiting microsatellite markers, fluorescent in situ hybridization (FISH) using repetitive DNA, inter retroelement amplified polymorphisms (IRAP) and some other molecular markers and morphological characters to estimate genetic diversity and relationships within and between different genus and species of Triticeae, specially Aegilops tauschii, (diploid, D genome) across Iran.
The diploid goatgrassAe. tauschii Coss. [Syn. Triticum tauschii (Coss.) Schmal., Aegilops squarrosa auct. non L., 2n=2x=14] contains the D genome of Triticeae cereals, and is considered the progenitor of the D genome in bread wheat (Triticum aestivum L., 2n=6x=42, genome composition AABBDD).
In our study on biodiversity of Aegilops tauschii in Iran using microsatellite markers, IRAP and FISH, all this markers revealed high level of polymorphism between populations of this species in Iran, but no clear groupings supporting the taxonomic groping were detected that can be a consequence of high gene flow between populations in this area.
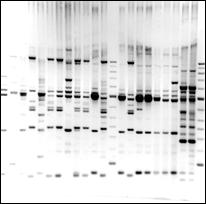 Polymorphism pattern from 20 Aegilops tauschii populations
generated by IRAP using Sukula and LTR6150
primers, showing high level of polymorphism between populations.
Polymorphism pattern from 20 Aegilops tauschii populations
generated by IRAP using Sukula and LTR6150
primers, showing high level of polymorphism between populations.
Ā
phh4(a)le.ac.uk
www.molcyt.com
January 2006